
Mixed-effects regression
Martijn Wieling (University of Groningen)
This lecture
- Introduction
- Recap: multiple regression
- Mixed-effects regression analysis: explanation
- Case-study: Influence of alcohol on L1 & L2
- Conclusion
Introduction
- Consider the following situation (taken from Clark, 1973):
- Mr. A and Mrs. B study reading latencies of verbs and nouns
- Each randomly selects 20 words and tests 50 subjects
- Mr. A finds (using a sign test) verbs to have faster responses
- Mrs. B finds nouns to have faster responses
- How is this possible?
The language-as-fixed-effect fallacy
- The problem is that Mr. A and Mrs. B disregard the (huge) variability in the words
- Mr. A included a difficult noun, but Mrs. B included a difficult verb
- Their set of words does not constitute the complete population of nouns and verbs, therefore their results are limited to their words
- This is known as the language-as-fixed-effect fallacy (LAFEF)
- Fixed-effect factors have repeatable and a small number of levels
- Word is a random-effect factor (a non-repeatable random sample from a larger population)
Why linguists are not always good statisticians
- LAFEF occurs frequently in linguistic research until the 1970’s
- Many reported significant results are wrong (the method is anti-conservative)!
- Clark (1973) combined a by-subject (\(F_1\)) analysis and by-item (\(F_2\)) analysis in measure min F’
- Results are significant and generalizable across subjects and items when min F’ is significant
- Unfortunately many researchers (>50%!) incorrectly interpreted this study and may report wrong results (Raaijmakers et al., 1999)
- E.g., they only use \(F_1\) and \(F_2\) and not min F’ or they use \(F_2\) while unneccesary (e.g., counterbalanced design)
Our problems solved…
- Apparently, analyzing this type of data is difficult…
- Fortunately, using mixed-effects regression models solves these problems!
- The method is easier than using the approach of Clark (1973)
- Results can be generalized across subjects and items
- Mixed-effects models are robust to missing data (Baayen, 2008, p. 266)
- We can easily test if it is necessary to treat item as a random effect
- No balanced design necessary (as in repeated-measures ANOVA)!
- But first some words about regression…
Recap: multiple regression (1)
- Multiple regression: predict one numerical variable on the basis of other independent variables (numerical or categorical)
- We can write a regression formula as \(y = \beta_0 + \beta_1x_1 + \beta_2x_2 + ... + \epsilon\)
- \(y\): (value of the) dependent variable
- \(x_i\): (value of the) predictor
- \(\beta_0\): intercept, value of \(y\) when all \(x_i = 0\)
- \(\beta_i\): slope, change in \(y\) when the value of \(x_i\) increases with 1
- \(\epsilon\): residuals, difference between observed values and predicted (fitted) values
Recap: multiple regression (2)
- Factorial predictors are (automatically) represented by binary-valued predictors: \(x_i = 0\) (reference level) or \(x_i = 1\) (alternative level)
- Factor with \(n\) levels: \(n-1\) binary predictors
- Interpretation of factorial \(\beta_i\): change from reference to alternative level
- Example of regression formula:
- Predict the reaction time of a subject on the basis of word frequency, word length, and subject age: RT = 200 - 5WF + 3WL + 10SA
Mixed-effects regression modeling: introduction
- Mixed-effects regression distinguishes fixed effects and random-effect factors
- Fixed effects:
- All numerical predictors
- Factorial predictors with a repeatable and small number of levels (e.g., Sex)
- Random-effect factors:
- Only factorial predictors!
- Levels are a non-repeatable random sample from a larger population
- Generally a large number of levels (e.g., Subject, Item)
What are random-effects factors?
- Random-effect factors are factors which are likely to introduce systematic variation (here: subject and item)
- Some subjects have a slow response (RT), while others are fast
= Random Intercept for Subject (i.e. \(\beta_0\) varies per subject) - Some items are easy to recognize, others hard
= Random Intercept for Item (i.e. \(\beta_0\) varies per item) - The effect of item frequency on RT might be higher for one subject than another: e.g., non-native participants might benefit more from frequent words than native participants
= Random Slope for Item Frequency per Subject (i.e. \(\beta_{\textrm{WF}}\) varies per subject) - The effect of subject age on RT might be different for one item than another: e.g., modern words might be recognized faster by younger participants
= Random Slope for Subject Age per Item (i.e. \(\beta_{\textrm{SA}}\) varies per item)
- Some subjects have a slow response (RT), while others are fast
- Note that it is essential to test for random slopes!
Question 1
Random slopes are necessary!
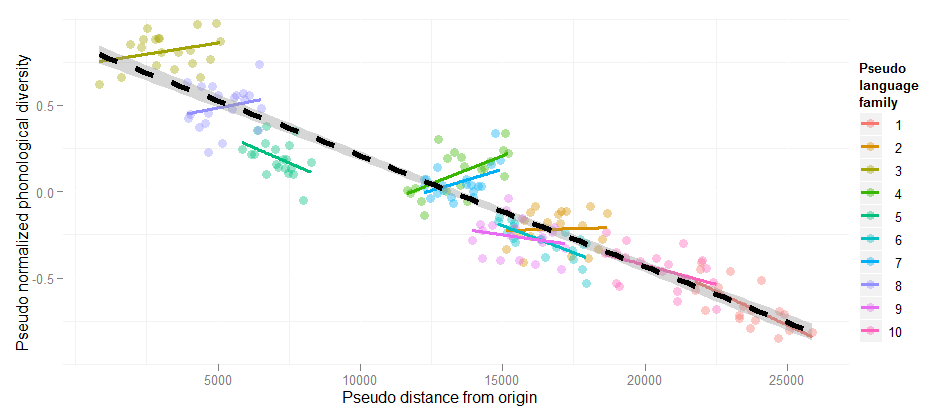
Estimate Std. Error t value Pr(>|t|)
Linear regression DistOrigin -6.418e-05 1.808e-06 -35.49 <2e-16
+ Random intercepts DistOrigin -2.224e-05 6.863e-06 -3.240 <0.001
+ Random slopes DistOrigin -1.478e-05 1.519e-05 -0.973 n.s.
(This example is explained at the HLP/Jaeger lab blog)
Modeling the variance structure
- Mixed-effects regression allow us to use random intercepts and slopes (i.e. adjustments to the population intercept and slopes) to include the variance structure in our data
- Parsimony: a single parameter (standard deviation) models this variation for every random slope or intercept (a normal distribution with mean 0 is assumed)
- The slope and intercept adjustments are Best Linear Unbiased Predictors
- Model comparison determines the inclusion of random intercepts and slopes
- Mixed-effects regression is only required when each level of the random-effect factor has multiple observations (e.g., participants respond to multiple items)
Specific models for every observation
- RT = 200 - 5WF + 3WL + 10SA (general model)
- The intercepts and slopes may vary (according to the estimated variation for each parameter) and this influences the word- and subject-specific values
- RT = 400 - 5WF + 3WL - 2SA (word: scythe)
- RT = 300 - 5WF + 3WL + 15SA (word: twitter)
- RT = 300 - 7WF + 3WL + 10SA (subject: non-native)
- RT = 150 - 5WF + 3WL + 10SA (subject: fast)
- And it is not hard to use!
lmer( RT ~ WF + WL + SA + (1+SA|Wrd) + (1+WF|Subj) )- (
lmerautomatically discovers random-effects structure: nested/crossed)
Variability in intercept and slope
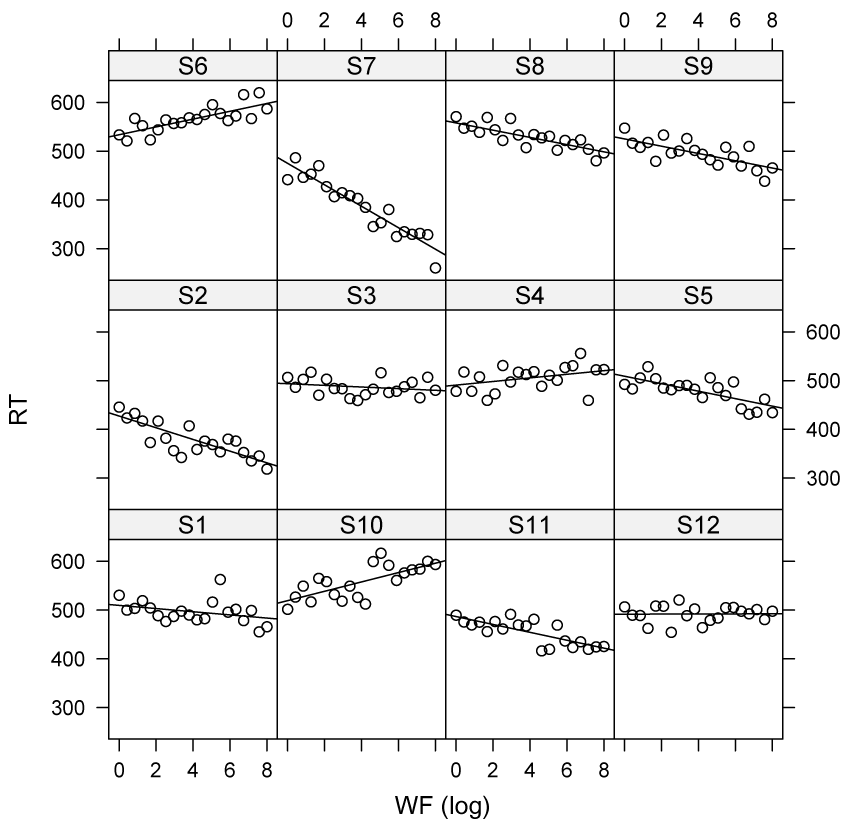
Random slopes and intercepts may be (cor)related
- For example:
| Subject | Intercept | WF slope |
|---|---|---|
| S1 | 525 | -2 |
| S2 | 400 | -1 |
| S3 | 500 | -2 |
| S4 | 550 | -3 |
| S5 | 450 | -2 |
| S6 | 600 | -4 |
| S7 | 300 | 0 |
BLUPs of lmer do not suffer from shrinkage
- The BLUPS (i.e. adjustment to the model estimates per item/participant) are close to the real adjustments, as
lmertakes into account regression towards the mean (fast subjects will be slower next time, and slow subjects will be faster) thereby avoiding overfitting and improving prediction - see Efron & Morris (1977)
Question 2
Center your variables (i.e. subtract the mean)!

- Otherwise random slopes and intercepts may show a spurious correlation
- Also helps the interpretation of interactions (see this lecture)
Mixed-effects regression assumptions
- Independent observations within each level of the random-effect factor
- Relation between dependent and independent variables linear
- No strong multicollinearity
- Residuals are not autocorrelated
- Homoscedasticity of variance in residuals
- Residuals are normally distributed
- (Similar assumptions as for regression)
Question 3
Model criticism
- Check the distribution of residuals
- If not normally distributed and/or heteroscedastic residuals then transform dependent variable or use generalized linear mixed-effects regression modeling
- Check outlier characteristics and refit the model when large outliers are excluded to verify that your effects are not ‘carried’ by these outliers
Model selection I
- “The data analyst knows more than the computer.” (Henderson & Velleman, 1981)
- Get to know your data!
- There is no adequate automatic procedure to find the best model
- Fitting the most complex random effects structure (Barr et al., 2013) is mostly not a good idea
- The model may not converge, or is uninterpretable (see Baayen et al., 2017)
- The power of your method is reduced (Matuschek et al., 2017)
Model selection II
- My stepwise variable-selection procedure (for exploratory analysis):
- Include random intercepts
- Add other potential explanatory variables one-by-one
- Insignificant predictors are dropped
- Test predictors for inclusion which were excluded at an earlier stage
- Test possible interactions (don’t make it too complex)
- Try to break the model by including significant predictors as random slopes
- Only choose a more complex model if supported by model comparison
Model selection III
- For a hypothesis-driven analysis, stepwise selection is problematic
- Confidence intervals too narrow: \(p\)-values too low (multiple comparisons)
- See, e.g., Burnham & Anderson (1998)
- Solutions:
- Careful specification of potential a priori models lining up with the hypotheses (including optimal random-effects structure) and evaluating only these models
- This may be followed by an exploratory procedure
- Validating a stepwise procedure via cross validation (e.g., bootstrap analysis)
- Careful specification of potential a priori models lining up with the hypotheses (including optimal random-effects structure) and evaluating only these models
Case study: influence of alcohol on L1 and L2
Case study: influence of alcohol on L1 and L2
- Reported in Wieling et al. (2019) and Offrede et al. (2020)
- Assess influence of alcohol (BAC) on L1 (clarity) vs. L2 (nativelikeness) ratings
- Prediction: higher BAC has negative effect on L1, but positive effect on L2 nativelikeness (based on Renner et al., 2018)
- ~80 recordings from native Dutch speakers included (all BACs < 0.8, no drugs)
- Dutch ratings were given by >100 native (sober) Dutch speakers (at Lowlands)
- English ratings were given by >100 native American English speakers (online)
- Dependent variable is \(z\)-transformed rating (5-point Likert scale)
- Numerical variables are centered (not \(z\)-transformed)
Dataset
SID BAC Sex BirthYear L2cnt SelfEN LivedEN L2anxiety Edu LID Lang Rating
1 S0045188-17 0.392 F -5.66 -1.04 0.463 Y 0.357 0.817 L0637009 NL 0.946
2 S0045188-17 0.392 F -5.66 -1.04 0.463 Y 0.357 0.817 L196 EN 0.330
3 S0045188-17 0.392 F -5.66 -1.04 0.463 Y 0.357 0.817 L86 EN -0.298
4 S0045188-17 0.392 F -5.66 -1.04 0.463 Y 0.357 0.817 L0614758 NL 0.325
5 S0045188-17 0.392 F -5.66 -1.04 0.463 Y 0.357 0.817 L220 EN -0.435
6 S0045188-17 0.392 F -5.66 -1.04 0.463 Y 0.357 0.817 L225 EN -0.239Fitting our first model
(fitted in R version 4.5.0 (2025-04-11 ucrt), lme4 version 1.1.37)
boundary (singular) fit: see help('isSingular') Estimate Std. Error t value
(Intercept) 0.144 0.0638 2.252
BAC -0.221 0.3262 -0.677 Groups Name Std.Dev.
LID (Intercept) 0.000
SID (Intercept) 0.560
Residual 0.738 Evaluating our hypothesis (1)
(note: random intercept for LID excluded)
- This model represents our hypothesis test (but likely without the correct random-effects structure)
Results (likely overestimating significance)
summary(m1, cor=F) # suppress estimated correlation of regression coefficient; Note: |t| > 2 => p < .05 (N > 100)Linear mixed model fit by REML ['lmerMod']
Formula: Rating ~ Lang * BAC + (1 | SID)
Data: lls
REML criterion at convergence: 5853
Scaled residuals:
Min 1Q Median 3Q Max
-3.668 -0.647 0.032 0.707 3.187
Random effects:
Groups Name Variance Std.Dev.
SID (Intercept) 0.305 0.552
Residual 0.533 0.730
Number of obs: 2541, groups: SID, 82
Fixed effects:
Estimate Std. Error t value
(Intercept) 0.0883 0.0635 1.39
LangNL 0.2430 0.0358 6.78
BAC -0.0592 0.3255 -0.18
LangNL:BAC -0.7286 0.1938 -3.76Visualization helps interpretation

- Interpretation: a higher BAC has a negative effect on L1, but not L2
Is the added interaction an improvement?
m0 <- lmer(Rating ~ Lang + (1|SID), data=lls) # comparison: best simpler model (BAC n.s.)
anova(m0, m1) # interaction necessaryData: lls
Models:
m0: Rating ~ Lang + (1 | SID)
m1: Rating ~ Lang * BAC + (1 | SID)
npar AIC BIC logLik -2*log(L) Chisq Df Pr(>Chisq)
m0 4 5866 5889 -2929 5858
m1 6 5855 5890 -2921 5843 14.7 2 0.00063 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Evaluating our hypothesis (2): adding a random slope
m2 <- lmer(Rating ~ Lang * BAC + (1 + Lang|SID), data=lls)
anova(m1, m2, refit=FALSE) # random slope necessary? (REML model comparison)Data: lls
Models:
m1: Rating ~ Lang * BAC + (1 | SID)
m2: Rating ~ Lang * BAC + (1 + Lang | SID)
npar AIC BIC logLik -2*log(L) Chisq Df Pr(>Chisq)
m1 6 5865 5900 -2927 5853
m2 8 5626 5672 -2805 5610 244 2 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1- This model represents the appropriate hypothesis test as it includes the adequate random-effects structure
Intermezzo: ML or REML model comparison?
- Restricted Maximum Likelihood (REML; default
lmerfitting method) is appropriate when comparing 2 models differing exclusively in the random-effects structure- The REML score depends on which fixed effects are in the model, so REML values are not comparable if the fixed effects change
- As REML is considered to give better estimates for the random effects, it is recommended to fit your final model (for reporting and inference) using REML
- ML is appropriate when comparing models differing only in the fixed effects
anovaautomatically refits the models with ML, unlessrefit=Fis specified
- Never compare 2 models differing in both fixed and random effects
- Only compare models which differ minimally (e.g., only in 1 random slope)
Results of hypothesis test
Estimate Std. Error t value
(Intercept) 0.0988 0.0747 1.322
LangNL 0.2544 0.0815 3.121
BAC 0.0994 0.3925 0.253
LangNL:BAC -0.9875 0.4265 -2.316- Note the higher |\(t\)|-values in the model without the random slope:
Random effects: correlation parameter
Groups Name Std.Dev. Corr
SID (Intercept) 0.653
LangNL 0.653 -0.83
Residual 0.683 - Note the absence of the random slope in the random-intercept-only model:
A note about random-effect correlation parameters
- In the previous model a correlation is assumed between random intercept and slope:
(1+X|RF)- Any terms before the
|are assumed to be correlated and a correlation parameter is calculated for each pair
- Any terms before the
- To exclude the correlation parameter, a simpler model can be specified as follows:
(1|RF) + (0+X|RF)(alternatively:(1+X||RF))- Model comparison (with
refit=FALSE) is used to determine the best model when only random effects differ
- Model comparison (with
- Factorial predictors (such as
Lang) should always be grouped with the intercept:(1+Lang|SID), but not:(1|SID) + (0 + Lang|SID)
Interpretation of correlation parameter
- Interpretation: speakers with higher ratings for English (intercept), have lower ratings for Dutch and vice versa (however: categorical variables are not centered)
Visualizing the results of the hypothesis test

- Interpretation as before: a higher BAC has a negative effect on L1, but not on L2
Alternative summary
m2b <- lmer(Rating ~ Lang : BAC + Lang + (1 + Lang|SID), data=lls) # instead of Lang * BAC
summary(m2b)$coef # effect of BAC per language separately Estimate Std. Error t value
(Intercept) 0.0988 0.0747 1.322
LangNL 0.2544 0.0815 3.121
LangEN:BAC 0.0994 0.3925 0.253
LangNL:BAC -0.8882 0.2684 -3.309 Estimate Std. Error t value
(Intercept) 0.0988 0.0747 1.322
LangNL 0.2544 0.0815 3.121
BAC 0.0994 0.3925 0.253
LangNL:BAC -0.9875 0.4265 -2.316Exploratory analysis
- Hypothesis seems partially supported
- But this may be caused by other variables:
- L2 anxiety
- L2 self rating
- (Other control variables)
- It may also be caused by the presence of atypical outliers
- Next step: exploratory analysis
Effect of speaker’s sex?
Estimate Std. Error t value
(Intercept) 0.1347 0.0870 1.549
LangNL 0.2532 0.0815 3.107
BAC 0.1325 0.3987 0.332
SexM -0.0815 0.0973 -0.838
LangNL:BAC -0.9801 0.4265 -2.298[1] 0.404Effect of speaker’s year of birth?
Estimate Std. Error t value
(Intercept) 0.09969 0.07472 1.334
LangNL 0.25462 0.08171 3.116
BAC 0.08313 0.39269 0.212
BirthYear 0.00603 0.00495 1.217
LangNL:BAC -0.99645 0.42764 -2.330[1] 0.217Effect of speaker’s education?
Estimate Std. Error t value
(Intercept) 0.102 0.0709 1.443
LangNL 0.255 0.0817 3.115
BAC 0.200 0.3734 0.536
Edu 0.112 0.0341 3.288
LangNL:BAC -1.034 0.4266 -2.424[1] 0.00139Effect of living in an English-speaking country?
Estimate Std. Error t value
(Intercept) 0.0831 0.0732 1.134
LangNL 0.2526 0.0817 3.091
BAC 0.2027 0.3708 0.546
Edu 0.1126 0.0342 3.290
LivedENY 0.1116 0.1147 0.973
LangNL:BAC -1.0368 0.4264 -2.432[1] 0.327Effect of self-rated English proficiency?
Estimate Std. Error t value
(Intercept) 0.1077 0.0644 1.671
LangNL 0.2491 0.0824 3.021
BAC 0.1675 0.3401 0.493
Edu 0.0758 0.0343 2.211
SelfEN 0.1299 0.0370 3.507
LangNL:BAC -1.0967 0.4283 -2.561[1] 0.00135Self-rated English proficiency: interaction?
Estimate Std. Error t value
(Intercept) 0.1087 0.0612 1.775
LangNL 0.2474 0.0753 3.285
BAC 0.0665 0.3236 0.206
Edu 0.0741 0.0344 2.157
SelfEN 0.2874 0.0541 5.309
LangNL:BAC -0.9385 0.3933 -2.386
LangNL:SelfEN -0.2519 0.0630 -3.996[1] 0.000103Visual interpretation of interaction

- Interpretation: a higher self-rated English proficiency is only predictive for the English ratings, not the Dutch ratings (unsurprisingly)
Effect of L2 count?
m6 <- lmer(Rating ~ Lang * BAC + Edu + SelfEN * Lang + L2cnt + (1 + Lang|SID), data=lls)
summary(m6)$coef Estimate Std. Error t value
(Intercept) 0.1080 0.0599 1.802
LangNL 0.2500 0.0755 3.314
BAC 0.1092 0.3172 0.344
Edu 0.0635 0.0339 1.875
SelfEN 0.2787 0.0531 5.246
L2cnt 0.1012 0.0467 2.168
LangNL:BAC -0.9356 0.3937 -2.377
LangNL:SelfEN -0.2499 0.0632 -3.956[1] 0.028- Note that the variable
Edunow does not have an absolute \(t\)-value > 2 anymore, but we retain it as it is close to significance (\(p \approx 0.06\) based onanova)
For both English and Dutch, or only English?
m7 <- lmer(Rating ~ Lang * BAC + Edu + SelfEN * Lang + L2cnt * Lang + (1 + Lang|SID), data=lls)
summary(m7)$coef Estimate Std. Error t value
(Intercept) 0.1076 0.0601 1.792
LangNL 0.2501 0.0758 3.300
BAC 0.1237 0.3183 0.389
Edu 0.0630 0.0338 1.863
SelfEN 0.2746 0.0536 5.127
L2cnt 0.1340 0.0677 1.979
LangNL:BAC -0.9640 0.3972 -2.427
LangNL:SelfEN -0.2432 0.0642 -3.785
LangNL:L2cnt -0.0576 0.0861 -0.668[1] 0.49Effect of L2 anxiety?
m7 <- lmer(Rating ~ Lang * BAC + Edu + SelfEN * Lang + L2cnt + L2anxiety + (1 + Lang|SID), data=lls)
summary(m7)$coef Estimate Std. Error t value
(Intercept) 0.1098 0.0580 1.892
LangNL 0.2453 0.0755 3.251
BAC 0.1042 0.3072 0.339
Edu 0.0606 0.0327 1.854
SelfEN 0.2026 0.0599 3.384
L2cnt 0.1202 0.0456 2.638
L2anxiety -0.2277 0.0918 -2.481
LangNL:BAC -0.9463 0.3931 -2.407
LangNL:SelfEN -0.2526 0.0632 -3.998[1] 0.0118For both English and Dutch, or only English?
m8 <- lmer(Rating ~ Lang * BAC + Edu + SelfEN * Lang + L2cnt + L2anxiety * Lang + (1 + Lang|SID), data=lls)
summary(m8)$coef Estimate Std. Error t value
(Intercept) 0.1103 0.0581 1.900
LangNL 0.2462 0.0756 3.259
BAC 0.0949 0.3075 0.309
Edu 0.0607 0.0327 1.857
SelfEN 0.1721 0.0680 2.531
L2cnt 0.1206 0.0456 2.644
L2anxiety -0.3202 0.1340 -2.389
LangNL:BAC -0.9293 0.3940 -2.359
LangNL:SelfEN -0.1973 0.0861 -2.292
LangNL:L2anxiety 0.1653 0.1747 0.946[1] 0.337Best model?
- Since SID is the only random-effect factor, and the characteristics are unique per SID, there are no additional random slopes to test
- All fixed-effects were tested (not all n.s. interactions were shown)
- Conclusion:
m7is the best model!
Testing assumptions: multicollinearity
Testing assumptions: autocorrelation
Testing assumptions: normality
Testing assumptions: heteroscedasticity
Model criticism: countering outlier influence
lls2 <- lls[ abs(scale(resid(m7))) < 2.5 , ] # trimmed model: 98.5% of original data
m7.2 <- lmer(Rating ~ Lang * BAC + Edu + SelfEN * Lang + L2cnt + L2anxiety + (1 + Lang|SID), data = lls2)
summary(m7.2)$coef Estimate Std. Error t value
(Intercept) 0.1161 0.0594 1.955
LangNL 0.2493 0.0758 3.289
BAC 0.1342 0.3145 0.427
Edu 0.0627 0.0332 1.887
SelfEN 0.2055 0.0612 3.360
L2cnt 0.1245 0.0463 2.687
L2anxiety -0.2451 0.0933 -2.626
LangNL:BAC -0.9850 0.3948 -2.495
LangNL:SelfEN -0.2708 0.0636 -4.260- The BAC interaction with language remains the same: significantly more negative effect of BAC on ratings for Dutch compared to English
Question 4
Checking the residuals of the trimmed model
Bootstrap sampling shows similar results
2.5 % 97.5 %
.sig01 0.42482 0.6031
.sig02 -0.88736 -0.6747
.sig03 0.49042 0.7369
.sigma 0.62306 0.6593
(Intercept) 0.00145 0.2273
LangNL 0.10500 0.4073
BAC -0.48284 0.7246
Edu 0.00206 0.1296
SelfEN 0.07526 0.3238
L2cnt 0.03867 0.2142
L2anxiety -0.43556 -0.0652
LangNL:BAC -1.73253 -0.2097
LangNL:SelfEN -0.40298 -0.1429- Note that the 95% CI of the variable
Edudoes not contain 0 and therefore indicates it is a significant predictor
Recap
- We have learned:
- How to interpret mixed-effects regression results
- How to use
lmerto conduct mixed-effects regression - How to include random intercepts and random slopes in
lmerand why these are essential when you have multiple responses per subject or item
- Associated lab session:
- https://www.let.rug.nl/wieling/Statistics/Mixed-Effects/lab
- Lab session contains additional information: how to do multiple comparisons, and using other optimizers (countering convergence problems)
- https://www.let.rug.nl/wieling/Statistics/Mixed-Effects/lab
Evaluation
Questions?
Thank you for your attention!
https://www.martijnwieling.nl
m.b.wieling@rug.nl




