

Analyzing EEG data using GAMs
Lecture 4 of advanced regression for linguists
Martijn Wieling
Computational Linguistics Research Group
This lecture
- Introduction
- ERPs to study gender violations
- Research question
- Design
- Methods:
Rcode - Results
- Discussion
ERPs to study grammatical gender violations
- A P600 (a positivity 'around' 600 ms. after stimulus onset) is sensitive to grammatical violations
- An N400 (a negativity 'around' 400 ms. after stimulus onset) is modulated by semantic context and lexical properties of a word
- The P600/N400 are found by comparing the incorrect sentences to the correct sentences
- Native speakers appear to show a P600 for gender violations
- But analyzed by averaging over items and over subjects!
This study
- In this study we are interested in how non-native speakers respond to gender violations (joint work with Nienke Meulman)
- Gender is very hard to learn for L2 learners
- Even though behaviorally L2 learners might show correct responses, the brain may reveal differences in processing gender
Research question
- Is the P600 for gender violations dependent on age of arrival for the L2 learners of German?
ERP data
- Today: analysis of single region of interest (ROI 8)
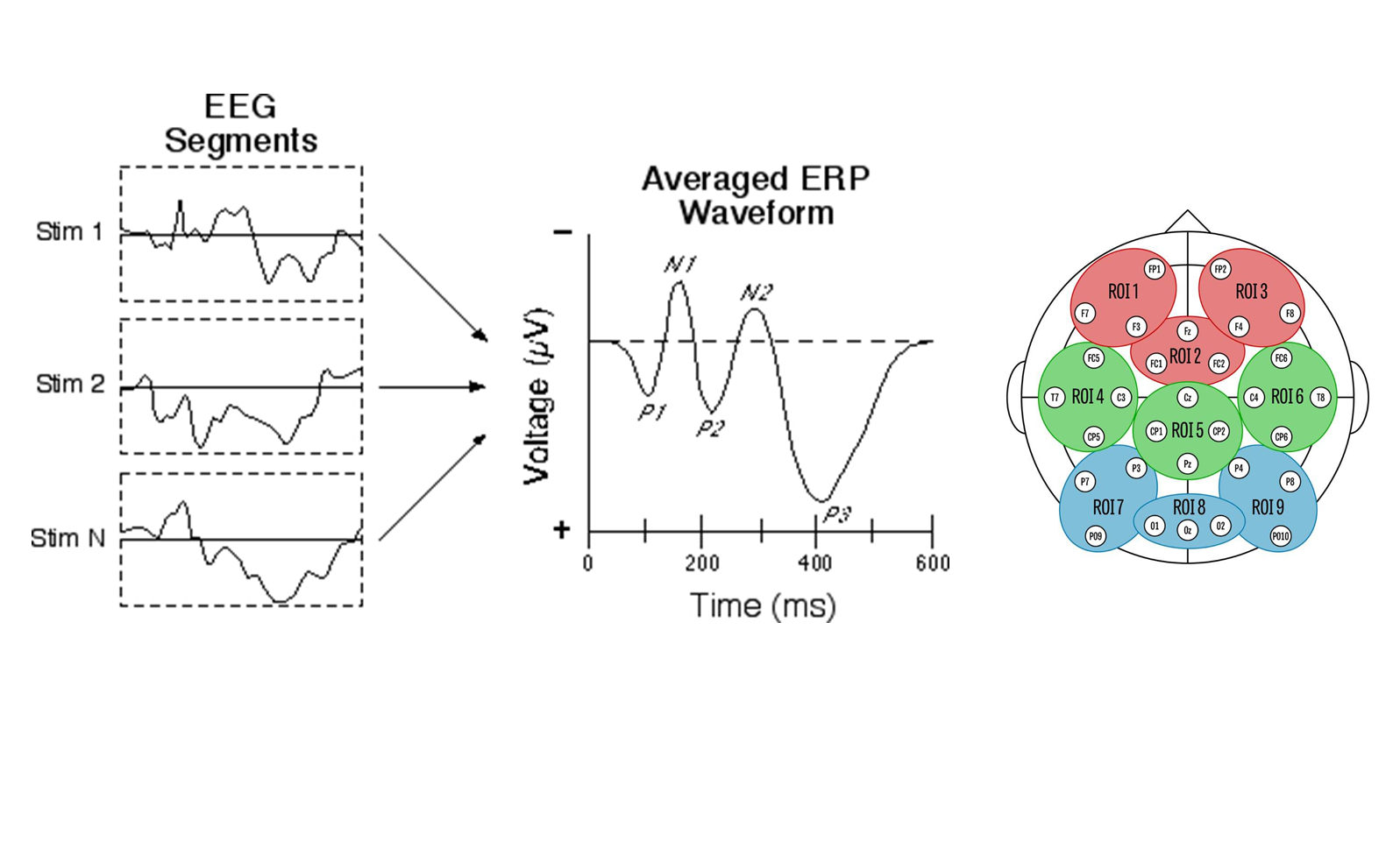
Design
- 67 L2 speakers of German (Slavic L1)
- Auditory presentation of correct sentences or sentences with a gender violation (incorrect determiner; no determiners in L1)
- 48 items in each condition: 96 trials per participant (minus artifacts)
- Example:
Nach der Schlägerei ist das/*der Auge des Angestellten von der Krankenschwester versorgt worden.
After the fight theneut/*themasc eye of the worker was treated by the nurse
Data overview
(R version 3.3.2 (2016-10-31), mgcv version 1.8.16, itsadug version 2.2.4)
library(mgcv)
library(itsadug)
load("dat.rda")
dat <- start_event(dat, event = c("Subject", "TrialNr")) # sorted data per subject and trial
head(dat)
# uV Time Subject Group Word TrialNr Roi Structure Correctness L1 AoArr LoR Age Edu
# 1 37.6 505 GL103 GenEarly Brot 1 post.mid DAN cor RU 8 15 23 3
# 2 38.1 515 GL103 GenEarly Brot 1 post.mid DAN cor RU 8 15 23 3
# 3 39.9 525 GL103 GenEarly Brot 1 post.mid DAN cor RU 8 15 23 3
# 4 28.4 535 GL103 GenEarly Brot 1 post.mid DAN cor RU 8 15 23 3
# 5 34.8 545 GL103 GenEarly Brot 1 post.mid DAN cor RU 8 15 23 3
# 6 42.4 555 GL103 GenEarly Brot 1 post.mid DAN cor RU 8 15 23 3
# AoAGroup start.event
# 1 Group 1: AoArr 7-12 TRUE
# 2 Group 1: AoArr 7-12 FALSE
# 3 Group 1: AoArr 7-12 FALSE
# 4 Group 1: AoArr 7-12 FALSE
# 5 Group 1: AoArr 7-12 FALSE
# 6 Group 1: AoArr 7-12 FALSE
dim(dat)
# [1] 442160 16
Much individual variation
(the signal has been downsampled to 100 Hz)
General patterns exist
(note the arbitrary age splits, however)
Question 1
Investigating difference between correct and incorrect
(for all participants; to limit CPU time, the time window was set to [500,1300])
dat$SubjectCor <- interaction(dat$Subject, dat$Correctness)
dat$WordCor <- interaction(dat$Word, dat$Correctness)
# duration discrete=F: 3000 sec.; 1/2/4/8/16 threads: 800/450/250/150/200 sec.
system.time(m0 <- bam(uV ~ s(Time, by = Correctness) + Correctness + s(Time, SubjectCor, bs = "fs", m = 1) +
s(Time, WordCor, bs = "fs", m = 1), data = dat, rho = rhoval, AR.start = dat$start.event, discrete = T,
nthreads = 2))
# user system elapsed
# 869.004 0.044 450.301
system.time(smry0 <- summary(m0)) # storing summary as it also takes some time to compute
# user system elapsed
# 75.777 0.004 75.830
Global difference between correct and incorrect
smry0 # show summary
#
# Family: gaussian
# Link function: identity
#
# Formula:
# uV ~ s(Time, by = Correctness) + Correctness + s(Time, SubjectCor,
# bs = "fs", m = 1) + s(Time, WordCor, bs = "fs", m = 1)
#
# Parametric coefficients:
# Estimate Std. Error t value Pr(>|t|)
# (Intercept) -0.553 0.473 -1.17 0.24
# Correctnessincor 0.733 0.670 1.09 0.27
#
# Approximate significance of smooth terms:
# edf Ref.df F p-value
# s(Time):Correctnesscor 1.04 1.08 0.39 0.57
# s(Time):Correctnessincor 3.32 4.32 6.76 1.2e-05 ***
# s(Time,SubjectCor) 114.28 1204.00 0.69 < 2e-16 ***
# s(Time,WordCor) 134.33 1726.00 0.28 < 2e-16 ***
# ---
# Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#
# R-sq.(adj) = 0.0513 Deviance explained = 5.18%
# fREML = 1.4922e+06 Scale est. = 281.02 n = 442160
Visualizing difference between correct and incorrect
par(mfrow = c(1, 2))
plot_smooth(m0, view = "Time", rug = F, plot_all = "Correctness", main = "", rm.ranef = T, eegAxis = T)
plot_diff(m0, view = "Time", comp = list(Correctness = c("incor", "cor")), rm.ranef = T, eegAxis = T)
Modeling the difference directly using a binary curve
dat$IsIncorrect <- (dat$Correctness == "incor") * 1 # creating a binary variable (incor: 1, cor: 0)
m0b <- bam(uV ~ s(Time) + s(Time, by = IsIncorrect) + s(Time, SubjectCor, bs = "fs", m = 1) + s(Time,
WordCor, bs = "fs", m = 1), data = dat, rho = rhoval, AR.start = dat$start.event, discrete = T, nthreads = 2)
# s(Time,by=IsIncorrect) = 0 when IsIncorrect = 0, otherwise it is a NON-CENTERED non-linear
# difference trajectory (i.e. including intercept difference)
smry0b <- summary(m0b)
smry0b$p.table
# Estimate Std. Error t value Pr(>|t|)
# (Intercept) -0.537 0.474 -1.13 0.257
smry0b$s.table # p(s(Time):IsIncorrect): significance of the full cor-incor DIFFERENCE
# edf Ref.df F p-value
# s(Time) 1.64 2.05 0.666 5.36e-01
# s(Time):IsIncorrect 4.08 5.00 3.833 1.80e-03
# s(Time,SubjectCor) 114.28 1204.00 0.688 1.46e-125
# s(Time,WordCor) 134.33 1726.00 0.275 7.19e-55
Modeling the difference using an ordered factor
dat$CorrectnessO <- as.ordered(dat$Correctness) # creating an ordered factor ...
contrasts(dat$CorrectnessO) <- "contr.treatment" # ... with contrast treatment
m0o <- bam(uV ~ s(Time) + s(Time, by = CorrectnessO) + CorrectnessO + s(Time, SubjectCor, bs = "fs",
m = 1) + s(Time, WordCor, bs = "fs", m = 1), data = dat, rho = rhoval, AR.start = dat$start.event,
discrete = T, nthreads = 2)
# s(Time,by=CorrecnessO) = 0 when CorrectnessO is at its reference level (cor), otherwise it is a
# CENTERED non-linear difference trajectory
smry0o <- summary(m0o)
smry0o$p.table # p(CorrectnessOincor) indicates significance of the constant DIFFERENCE
# Estimate Std. Error t value Pr(>|t|)
# (Intercept) -0.537 0.474 -1.13 0.257
# CorrectnessOincor 0.753 0.670 1.12 0.261
smry0o$s.table # p(s(Time):CorrectnessOincor) indicates significance of non-linear DIFFERENCE
# edf Ref.df F p-value
# s(Time) 1.64 2.05 0.666 5.36e-01
# s(Time):CorrectnessOincor 3.08 4.00 4.578 1.08e-03
# s(Time,SubjectCor) 114.28 1204.00 0.688 1.47e-125
# s(Time,WordCor) 134.33 1726.00 0.275 7.19e-55
Visualization of both difference curves
par(mfrow = c(1, 2))
plot(m0b, select = 2, shade = T, rug = F, main = "Binary difference curve", ylim = c(3, -3))
plot(m0o, select = 2, shade = T, rug = F, main = "Ordered factor difference curve", ylim = c(3, -3))
Question 2
Testing our research question: a non-linear interaction
(te is used to model a non-linear interaction with predictors on a different scale)
m1 <- bam(uV ~ te(Time, AoArr, by = Correctness) + Correctness + s(Time, SubjectCor, bs = "fs", m = 1) +
s(Time, WordCor, bs = "fs", m = 1), data = dat, rho = rhoval, AR.start = dat$start.event, discrete = T,
nthreads = 2)
smry1 <- summary(m1)
smry1$p.table
# Estimate Std. Error t value Pr(>|t|)
# (Intercept) -0.489 0.469 -1.041 0.298
# Correctnessincor 0.595 0.665 0.895 0.371
smry1$s.table
# edf Ref.df F p-value
# te(Time,AoArr):Correctnesscor 3.10 3.19 1.680 1.52e-01
# te(Time,AoArr):Correctnessincor 5.88 6.95 4.912 1.60e-05
# s(Time,SubjectCor) 111.94 1202.00 0.659 2.24e-119
# s(Time,WordCor) 134.38 1726.00 0.276 5.93e-55
Visualization of the 2-dimensional difference
Note the default maximum number of edf's per smooth: 24 (5\(^2\) - 1)
plot_diff2(m1, view = c("Time", "AoArr"), rm.ranef = T, comp = list(Correctness = c("incor", "cor")))
fadeRug(dat$Time, dat$AoArr) # indicate points without data
Interpreting the 2-dimensional interaction
(note that the \(y\)-axes are not inverted here)
Interpret interactions yourself!
Decomposition: the pure effect of age of arrival
m2 <- bam(uV ~ s(Time, by = Correctness) + s(AoArr, by = Correctness) + ti(Time, AoArr, by = Correctness) +
Correctness + s(Time, SubjectCor, bs = "fs", m = 1) + s(Time, WordCor, bs = "fs", m = 1), data = dat,
rho = rhoval, AR.start = dat$start.event, discrete = T, nthreads = 2)
smry2 <- summary(m2)
smry2$p.table
# Estimate Std. Error t value Pr(>|t|)
# (Intercept) -0.478 0.470 -1.02 0.308
# Correctnessincor 0.579 0.665 0.87 0.384
smry2$s.table
# edf Ref.df F p-value
# s(Time):Correctnesscor 1.04 1.09 0.158 7.40e-01
# s(Time):Correctnessincor 3.32 4.31 6.756 1.27e-05
# s(AoArr):Correctnesscor 1.03 1.04 2.588 1.10e-01
# s(AoArr):Correctnessincor 1.05 1.05 3.714 4.76e-02
# ti(Time,AoArr):Correctnesscor 1.06 1.11 2.058 1.33e-01
# ti(Time,AoArr):Correctnessincor 1.93 2.64 0.336 7.08e-01
# s(Time,SubjectCor) 111.86 1202.00 0.654 7.14e-119
# s(Time,WordCor) 134.38 1726.00 0.276 5.94e-55
A simpler model without the non-linear interaction
m3 <- bam(uV ~ s(Time, by = Correctness) + s(AoArr, by = Correctness) + Correctness + s(Time, SubjectCor,
bs = "fs", m = 1) + s(Time, WordCor, bs = "fs", m = 1), data = dat, rho = rhoval, AR.start = dat$start.event,
discrete = T, nthreads = 2)
smry3 <- summary(m3)
smry3$p.table
# Estimate Std. Error t value Pr(>|t|)
# (Intercept) -0.475 0.470 -1.011 0.312
# Correctnessincor 0.582 0.666 0.873 0.383
smry3$s.table
# edf Ref.df F p-value
# s(Time):Correctnesscor 1.01 1.03 0.375 5.54e-01
# s(Time):Correctnessincor 3.32 4.32 6.762 1.24e-05
# s(AoArr):Correctnesscor 1.05 1.06 2.705 1.04e-01
# s(AoArr):Correctnessincor 1.11 1.12 3.337 5.28e-02
# s(Time,SubjectCor) 111.78 1202.00 0.652 1.83e-118
# s(Time,WordCor) 134.38 1726.00 0.276 5.96e-55
Model comparison
(workaround to use fREML comparison with discrete=T; select=T treats smooths as random effects)
m2.alt <- bam(uV ~ s(Time, by = Correctness) + s(AoArr, by = Correctness) + ti(Time, AoArr, by = Correctness) +
Correctness + s(Time, SubjectCor, bs = "fs", m = 1) + s(Time, WordCor, bs = "fs", m = 1), data = dat,
rho = rhoval, AR.start = dat$start.event, method = "fREML", select = T, discrete = T, nthreads = 2)
m3.alt <- bam(uV ~ s(Time, by = Correctness) + s(AoArr, by = Correctness) + Correctness + s(Time, SubjectCor,
bs = "fs", m = 1) + s(Time, WordCor, bs = "fs", m = 1), data = dat, rho = rhoval, AR.start = dat$start.event,
method = "fREML", select = T, discrete = T, nthreads = 2)
compareML(m2.alt, m3.alt)
# m2.alt: uV ~ s(Time, by = Correctness) + s(AoArr, by = Correctness) +
# ti(Time, AoArr, by = Correctness) + Correctness + s(Time,
# SubjectCor, bs = "fs", m = 1) + s(Time, WordCor, bs = "fs",
# m = 1)
#
# m3.alt: uV ~ s(Time, by = Correctness) + s(AoArr, by = Correctness) +
# Correctness + s(Time, SubjectCor, bs = "fs", m = 1) + s(Time,
# WordCor, bs = "fs", m = 1)
#
# Chi-square test of fREML scores
# -----
# Model Score Edf Chisq Df p.value Sig.
# 1 m3.alt 1492233 14
# 2 m2.alt 1492233 20 0.528 6.000 0.983
#
# AIC difference: 5.95, model m3.alt has lower AIC.
Question 3
Difference curves
par(mfrow = c(1, 2))
plot_diff(m3, view = "Time", comp = list(Correctness = c("incor", "cor")), rm.ranef = T, eegAxis = T)
plot_diff(m3, view = "AoArr", comp = list(Correctness = c("incor", "cor")), rm.ranef = T, eegAxis = T)
Finally: model criticism
par(mfrow = c(1, 2))
qqp(resid_gam(m3)) # resid_gam takes autocorrelation into account
hist(resid_gam(m3))

Problematic residuals!
- This type of residual distribution is common for EEG data
- These deviations are problematic and may affect \(p\)-values
Approach to normalize residuals
- Model fitting using scaled-\(t\) distribution
- Implemented in
gam:family="scat" - Adding
rhotogampossible via custom function (created by Natalya Pya) - Unfortunately,
gamis too slow, so abamapproach is in development - Current state: implemented for
fREMLestimation with pre-specified scaled-\(t\) parameters (\(\theta\)) (see Meulman et al., 2015) - Thus
gamstill necessary to determine (\(\theta\)) (but applied to subset / simpler model specification)
- Implemented in
Estimating scaled-\(t\) parameters efficiently
- Fit an intercept-only scaled-t
gammodel (includingrho,method="REML") to all data and extract \(\theta\) - Fit the complete model without random effects as a scaled-\(t\)
gammodel (includingrho,method="REML") to all data and extract \(\theta\) - Compare the models' \(\theta\) values:
- Small differences: use \(\theta\) of the complex model in the
bamestimation - Large differences: determine \(\theta\) by fittting the complete model specification (including random effects) to a small subset of the data (randomly select trials, compare \(\theta\) across a few runs)
- Small differences: use \(\theta\) of the complex model in the
Fitting a scaled-\(t\) model
source("bam.art.fit-v11.R") # custom functions by Natalya Pya
g0 <- gam(uV ~ 1, data = dat, method = "REML", family = scat(rho = rhoval, AR.start = dat$start.event)) # takes 6 seconds
g1 <- gam(uV ~ s(Time, by = Correctness) + s(AoArr, by = Correctness) + Correctness, data = dat, method = "REML",
family = scat(rho = rhoval, AR.start = dat$start.event)) # takes 14 minutes
theta0 <- g0$family$getTheta(TRUE)
(theta1 <- g1$family$getTheta(TRUE))
# [1] 4.57 5.27
theta0 - theta1
# [1] 0.00275 0.00210
# the Gaussian model (m3) takes 250 sec. using 4 procs
system.time(m4.scat <- bam.art(uV ~ s(Time, by = Correctness) + s(AoArr, by = Correctness) + Correctness +
s(Time, SubjectCor, bs = "fs", m = 1) + s(Time, WordCor, bs = "fs", m = 1), data = dat, method = "fREML",
family = art(theta = theta1, rho = rhoval), AR.start = dat$start.event, discrete = T, nthreads = 4))
# user system elapsed
# 13213.34 7.68 3447.88
Using the scaled-\(t\) distribution: \(p\)-values change
summary(m3)$s.table
# edf Ref.df F p-value
# s(Time):Correctnesscor 1.01 1.03 0.375 5.54e-01
# s(Time):Correctnessincor 3.32 4.32 6.762 1.24e-05
# s(AoArr):Correctnesscor 1.05 1.06 2.705 1.04e-01
# s(AoArr):Correctnessincor 1.11 1.12 3.337 5.28e-02
# s(Time,SubjectCor) 111.78 1202.00 0.652 1.83e-118
# s(Time,WordCor) 134.38 1726.00 0.276 5.96e-55
summary(m4.scat)$s.table
# edf Ref.df F p-value
# s(Time):Correctnesscor 1.03 1.05 0.674 4.27e-01
# s(Time):Correctnessincor 4.17 5.38 14.267 1.40e-14
# s(AoArr):Correctnesscor 1.48 1.50 1.802 2.74e-01
# s(AoArr):Correctnessincor 1.00 1.00 6.915 8.55e-03
# s(Time,SubjectCor) 120.61 1204.00 1.453 1.25e-310
# s(Time,WordCor) 137.74 1727.00 0.301 4.44e-62
Using the scaled-\(t\) distribution: similar patterns
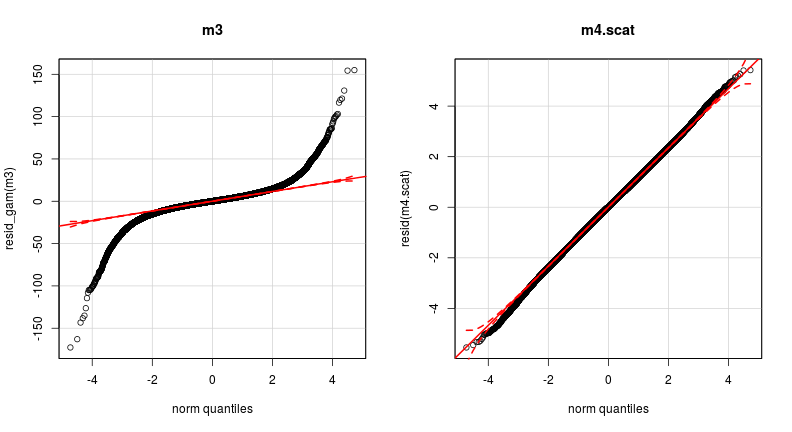
Model criticism: much improved!
par(mfrow = c(1, 2))
qqp(resid_gam(m3), main = "m3")
qqp(resid(m4.scat), main = "m4.scat") # resid of scat model takes autocorrelation into account
Discussion
- Still work to do:
- E.g., testing the significance of other possibly important variables (proficiency, age, etc.)
- But don't make it too complex!
- There is much variation present in EEG data and adding very complex surfaces will almost certainly improve your model significantly
- Keep it simple, otherwise you won't be able to compute/interpret the results
- Also note that for hypothesis testing, it is essential not to conduct step-wise approaches
Conclusion
- GAMs are very useful to analyze EEG and other time-series data
- The method is very suitable to detect non-linear patterns, while taking into account individual variation and correcting for autocorrelation
- The results of this study are published in PLOS ONE
- Supplementary material, including code and results is available here
Recap
- We have applied GAMs to EEG data and learned how to:
- model difference smooths directly using binary predictors and ordered factors
- use
te(Time,AoArr)to model a non-linear interaction - decompose
te(Time,AoArr)usingti()ands() - use a scaled-\(t\) distribution to improve residuals
- (fitting this type of model for a complete data set may take a lot of CPU time, however)
- While we have analyzed a single region of interest, GAMs allow for spatial distribution analyses
- E.g., via
te(x, y, Time, d = c(2,1)) - See http://eolomea.let.rug.nl/GAM/ScalpEEG
- E.g., via
- After the break:
- Next lecture: generalized additive logistic modeling using dialect data
Evaluation
Questions?
Thank you for your attention!